Addressing the challenge of antimicrobial resistance using innovative data approaches
Ashoka Faculty Fellows have pioneered a novel approach to studying AMR, surpassing the conventional exploration of individual drug-bug combinations
Antimicrobial resistance (AMR) is a complex and interdisciplinary public health challenge at a global level. Seven lakh people lose their lives every year to the disease that is caused by antibiotic-resistant pathogens. This number is projected to increase up to one crore by 2050 if we do not intervene. Over the past few years, numerous international organisations and national governments have launched initiatives to tackle the challenge of AMR. We now have identified many important factors that play a role in the AMR crisis. For instance, the increased use of antibiotics in humans, animals and agriculture increases the evolution of drug resistance. Similarly, reduced usage of prophylactic measures, such as vaccines, results in the increased usage of antibiotics and in turn increases AMR.
Widespread surveillance is essential across healthcare centres, agricultural sectors, animal farms, and wastewater for effectively combating AMR. However, AMR surveillance and studies traditionally focus on specific drug-bug combinations, like carbapenem-resistant Klebsiella pneumoniae.
Carbapenems serve as the last-resort antibiotics, reserved for treating infections that do not respond to more commonly used antibiotics. Klebsiella pneumoniae, a bacteria resistant to most known antibiotics, has become a significant concern for healthcare workers. This resistance poses a heightened threat to patients with advanced age, compromised immune systems, or prolonged hospitalisation, making them particularly susceptible to such infections.
While wastewater and environmental surveillance aims to detect Klebsiella and genes conferring carbapenem resistance, it is crucial to note that genes for resistance to one antibiotic often coexist with those for resistance to others. This complex relationship is also mirrored in their resistance profiles, where resistance to multiple antibiotics is commonly observed together.
To address these complexities, Dr Shraddha Karve and Dr Rintu Kutum, Faculty Fellows at the Trivedi School of Biosciences and the Department of Computer Sciences, respectively, proposed a novel analysis approach using Klebsiella as a proof of concept. They looked at the resistance profile of Klebsiella for a set of common antibiotics across publicly available global surveillance data and grouped all the possible resistance profiles as a ‘subtype’ from the data.
A resistance profile is a summary that shows how a microorganism, such as a bacterium, responds to different antibiotics. It indicates which antibiotics are effective in treating the microorganism and which ones it has developed resistance to. This information is essential for guiding healthcare decisions, helping to choose the most effective treatments and understanding patterns of antibiotic resistance in populations.
The researchers then looked at the predominant subtypes across different geographical regions and time (monthly). They further checked how different climatic variables, such as precipitation and temperature, correlate with the abundance of individual subtypes.
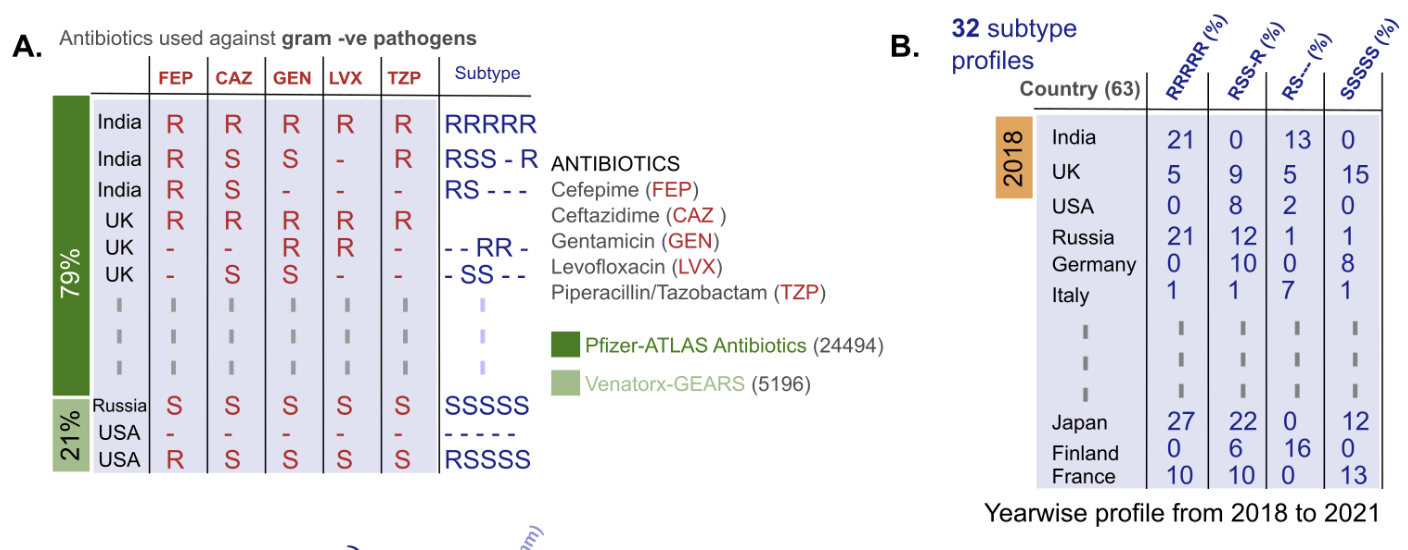
As a result of this comprehensive study done by Ashoka faculty fellows, it is shown that only a couple of subtypes dominate the landscape. Two subtypes, one that is sensitive to all the antibiotics and the other that is resistant to all antibiotics are predominant. Intermediate subtypes that are sensitive to a few antibiotics and resistant to others are not that common. This result reflects that resistance to different antibiotics goes hand in hand. Genes that make the bacteria resistant are also likely to be linked very tightly.
These results call for a holistic approach to studying AMR beyond a single drug-bug combination. It underlines the need to take into consideration the entire resistance profile of a bug for effective surveillance, diagnosis, and treatment.
This idea was proposed as a part of the ‘Global AMR Data Challenge’. The team, comprising four members from Ashoka—Dr Shraddha Karve, Dr Rintu Kutum, Vasundhara Karthikeyan and Ragul N (ASP’23, CS), alongside a collaborator Dr Debojit Kumar Sarma, Scientist-C, from ICMR-National Institute of Research in Environmental Health, secured the prestigious innovation award for their groundbreaking idea.
Approach: To accomplish this goal a custom-made computational workflow (please see the GitHub page for details) was used. Briefly, those five antibiotics were shortlisted that were available in public datasets# and are used to treat Klebsiella infections. The researchers then generated all the possible ‘subtypes’ i.e. a combined resistance profile, in the dataset. For example, the Klebsiella isolate that is resistant to all five antibiotics will be under the subtype RRRRR (where ‘R’ stands for ‘resistant’) and the Klebsiella isolate that is sensitive to the first antibiotic but resistant to the rest will be under the subtype SRRRR (where ‘S’ stands for sensitive). Over ten thousand isolates of Klebsiella that belonged to ~30 different subtype categories, were analysed.
(Edited by Dr Yukti Arora)